The Challenges of Precision Medicine and New Advances in Molecular Diagnostic Testing in Hematolymphoid Malignancies: Impact on the VHA
The Hematopathology Molecular Genetics subcommittee presents recommendations for molecular diagnostic testing in acute myeloid leukemia, myeloproliferative neoplasms, myelodysplastic syndrome, and lymphomas and for the development of an interfacility consultation service.
In January 2015, President Obama introduced the Precision Medicine Initiative, a program set up to identify new biomedical discoveries for the development of a personalized knowledge base of disease entities and individualized treatments. Advances in precision medicine typically involve the use of targeted therapies tailored to individual genetic characteristics identified with molecular testing. The goals are to improve survival and reduce adverse effects. With an initial budget of $215 million, this initiative presented a unique opportunity to combine efforts in genomic discovery, bioinformatic analysis, and health information technology to move toward data-driven, evidence-based precision medicine.1
The VHA is the largest comprehensive health care system in the U. S. and has more than 1,700 care sites serving nearly 9 million veterans each year. The budget for this single-payer system is proposed by the President and approved by Congress. As the VHA must treat a diverse and aging veteran population in an environment of rising costs and budget constraints, limited resources must be monitored and appropriated for the most cost-effective health care delivery. Precision medicine offers a model in which physicians can select the most appropriate diagnostic tests in defined clinical settings to direct clinical care. It supports the testing needed to subdivide each disease category into distinct subcategories. Nevertheless, the need for fiscal responsibility in a capitated health care system recommends testing in cases in which it can change therapy or prognosis rather than for purely academic reasons.
Pathology and Laboratory Medicine Service
Given limited resources and an increasing number of requests for advanced molecular testing, the VA Pathology and Laboratory Medicine Service (P&LMS) formed the Molecular Genetics Pathology Workgroup (MGPW) in September 2013. The charter listed the tasks of the MGPW to “provide recommendations on how to effectively use molecular genetics tests, promote increased quality and availability of testing within the VHA, encourage internal referral testing, provide an organizational structure for Molecular Genetics Testing Consortia, and create a P&LMS policy for molecular genetic testing in general, specifically addressing the issues surrounding laboratory developed testing.” The MGPW has 4 subcommittees: molecular oncology, pharmacogenetics, hematopathology molecular genetics (HMG), and genetic medicine. Since its inception, the HMG subcommittee has had several objectives:
- Standardize the molecular testing nomenclature for and develop practice guidelines for acute myeloid leukemia (AML), myeloproliferative neoplasms (MPN), myelodysplastic syndrome (MDS), chronic lymphocytic leukemia (CLL)/small lymphocytic lymphoma, lymphoma, and plasma cell neoplasms;
- Develop standardized reporting guidelines for current VA molecular laboratories;
- Identify new tests as they are being reported in the literature and collaborate with hematology and oncology services to evaluate the clinical utility of these tests for VA patients;
- Network current VA molecular laboratories, perform fact-finding for these laboratories, and compile test menus; and
- Assess for the formation of VA-wide interfacility consultation services for hematopathology so that all VA facilities, regardless of their complexity, will be able to access the expertise of hematopathology-trained pathologists (Appendix).
The HMG subcommittee met monthly and discussed various diagnostic entities in hematopathology. For hematolymphoid malignancies, it was generally agreed that the traditional laboratory tools of morphology, flow cytometry, and immunohistochemistry (IHC) are standard in initial assessment and often in diagnosis. As the clinical molecular and cytogenetic assays of karyotype, fluorescence in situ hybridization (FISH), advanced DNA sequencing, microarray, and highly sensitive polymerase chain reaction (PCR) analysis affect diagnosis, subclassification, minimal residual disease (MRD) monitoring, prognosis, and therapy selection, their use is marked by a high degree of variability. As a result, standardization is needed. As each laboratory develops and reports ancillary testing, the variable reporting formats may generate postanalytic errors.
A detailed description of all molecular methodologies is beyond the scope of this article. For practicing pathologists, challenges remain in overall cost and reimbursement, extensive and time-consuming data analysis, and in some cases, interpretation differences.
Myeloid Neoplasms
Myeloid malignancies were divided into AML, MPN, and MDS. Next-generation sequencing (NGS) information for these malignancies was used to identify various contributory functional categories, including cell signaling (FLT3, KIT, JAK2, MPL, KRAS/NRAS, PTPN11, NF1, CSF3R); transcription (CEBPA, RUNX1, GATA1/GATA2, PHF6, ETV6); splicing (SF3B1, SRSF2, ZRSR2, U2AF1); epigenetics (DNMT3A, TET2, IDH1/IDH2, ASXL1, EZH2, SUZ12, KDM6A); cohesin complex (STAG2, SMC1A, SMC3, RAD21); and cell cycle (TP53, NPM1).2
Acute Myeloid Leukemia
The HMG subcommittee reviewed the literature on prognostically significant genes in myeloid leukemias. Karyotype abnormalities, such as t(8;21) and inv(16), collectively known as the core-binding factor (CBF) leukemias, t(15;17), t(11q23) (KMT2A/MLL), and so forth, are recurrent lesions in AML. Included in the minimum set of genes recommended by the National Comprehensive Cancer Network (NCCN) for AML prognosis evaluation are nucleolar protein nucleophosmin (NPM1), CCAAT/enhancer-binding protein
Some of the chromosomal translocations, such as inv(16)/t(16;16) in AML and t(15;17) in acute promyelocytic leukemia, can be monitored with FISH or reverse transcription–PCR (RT-PCR) analysis. As NPM1 mutations tend to be seen in recurrence, they can be used as molecular markers for MRD. Other mutations that provide important prognostic information in AML include:
- Activating insertions/duplications in the FLT3 receptor tyrosine kinase, which can be detected with PCR sizing assays;
- Mutations in the KIT receptor tyrosine kinase, which can be detected with DNA sequencing or more limited hotspot PCR;
- Mutations in the DNA methyltransferase, DNMT3A, a poor prognostic indicator seen in 22% of cases of AML, also detected with gene sequencing or more limited hotspot PCR; and
- Another set of genes, TET2, IDH1, IDH2, KRAS, NRAS, EZH2, and ASXL1, is mutated in MPN as well as AML and MDS, making a common molecular panel with next-generation sequencing useful in diagnosing and risk-stratifying all myeloid neoplasms.
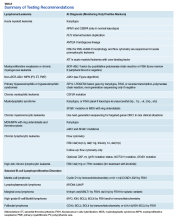
The HMG subcommittee agreed that, for de novo AML, chromosomal karyotype is the standard of care, necessary in detecting known cytogenetic abnormalities as well as a wide range of lesions that might indicate a diagnosis of AML with myelodysplasia-related changes at time of diagnosis. In addition, molecular analysis of FLT3 is useful in determining prognosis, and CEBPA (biallelic) and NPM1 mutations are good prognostic factors in normal-karyotype AML. KMT2A (MLL) rearrangements should be tested with FISH if the lineage is ambiguous. The PML-RARA fusion gene also should be tested with FISH if morphologic and flow cytometry results suggest acute promyelocytic leukemia (Table). At this time, testing for TP53, DNMT3A, RAS, and other such mutations is not recommended because it is not cost-effective for the VA.
Myeloproliferative Neoplasms
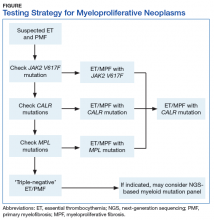
Myeloproliferative neoplasms are clonal hematopoietic stem cell disorders characterized by proliferation of at least 1 myeloid lineage: granulocytic, erythroid, or megakaryocytic. Myeloproliferative neoplasms show a range of recurrent chromosomal translocations, such as BCR-ABL1 fusion in chronic myelogenous leukemia (CML) that can be detected with RT-PCR analysis as well as FISH. In CML, BCR-ABL1 fusion transcript levels detected by a quantitative PCR (qPCR) method are now used to monitor the course of CML therapy with tyrosine kinase inhibitors (TKIs) and to trigger a treatment change in drug-resistant cases. Given the importance of qPCR in clinical management, significant progress has been made in standardizing both the PCR protocol and the reference materials used to calibrate the BCR-ABL1 PCR assay. BCR-ABL1–negative MPN, including polycythemia vera (PV), essential thrombocythemia (ET), and primary myelofibrosis (PMF), are most commonly associated with mutations in the tyrosine kinase JAK2. Mutations in CALR and MPL are seen in a subset of patients with ET and PMF as well, whereas PV is essentially exclusively a disease of JAK2 mutations.
Chronic myelogenous leukemia is the prototypical MPN. To establish the initial diagnosis, FISH and/or qPCR for BCR-ABL1 fusion should be used. If CML is confirmed, the sample can be reflexed to qPCR BCR-ABL1 on the initial peripheral blood and/or bone marrow sample(s) to establish the patient’s baseline. In addition, a bone marrow sample (aspirate) should be used for a complete karyotype and for morphologic confirmation of disease phase.